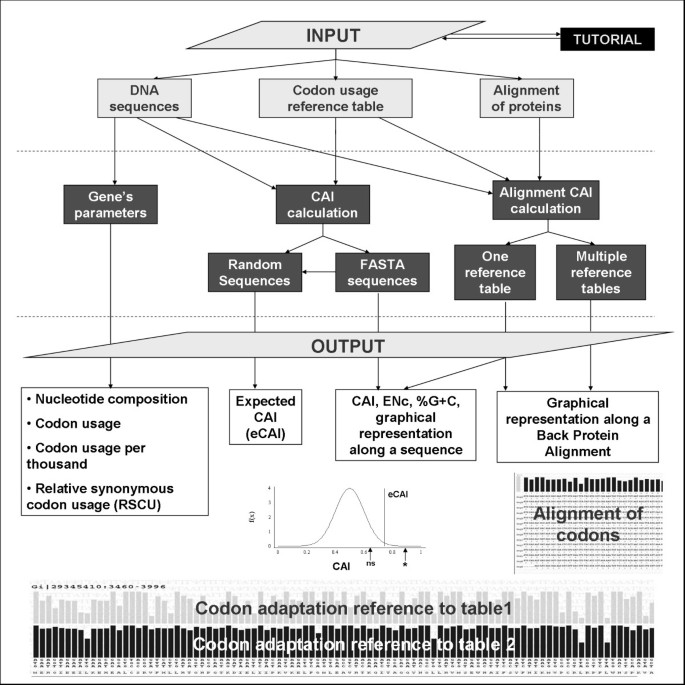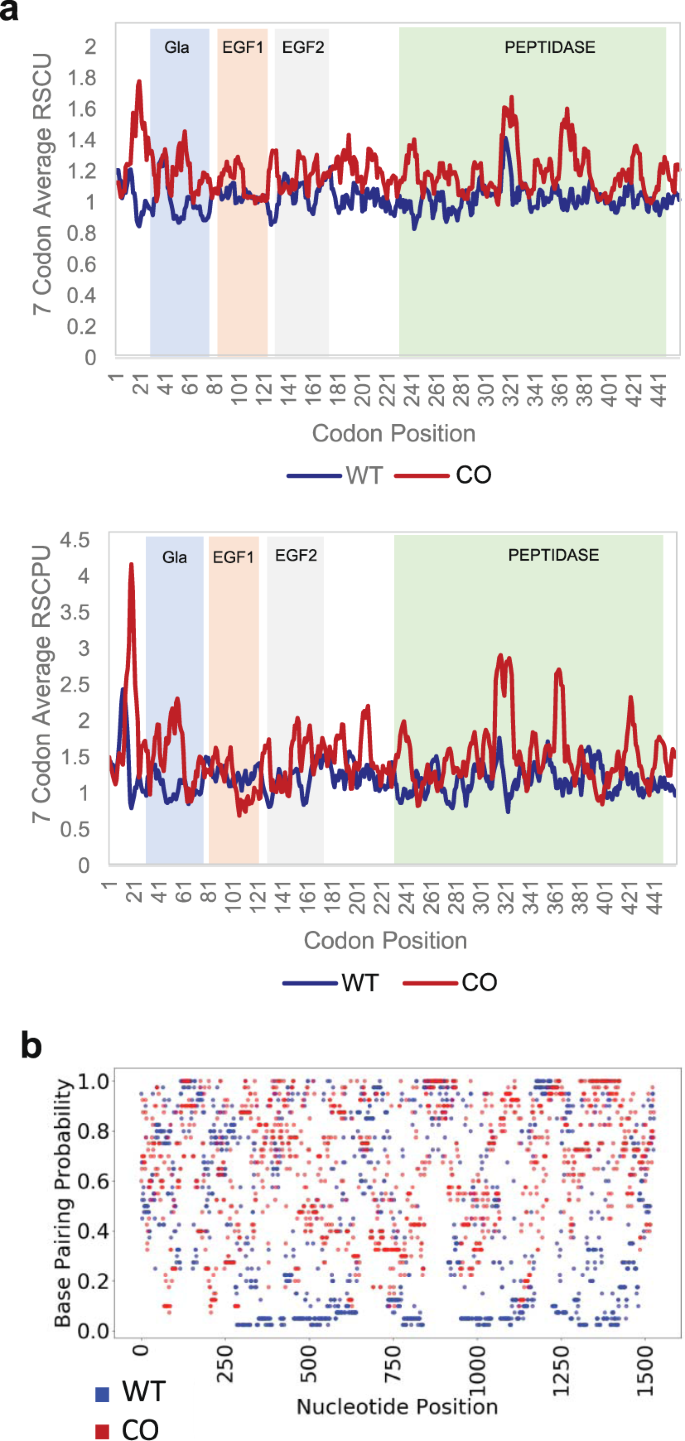
Effects of codon optimization on coagulation factor IX translation and structure: Implications for protein and gene therapies | Scientific Reports
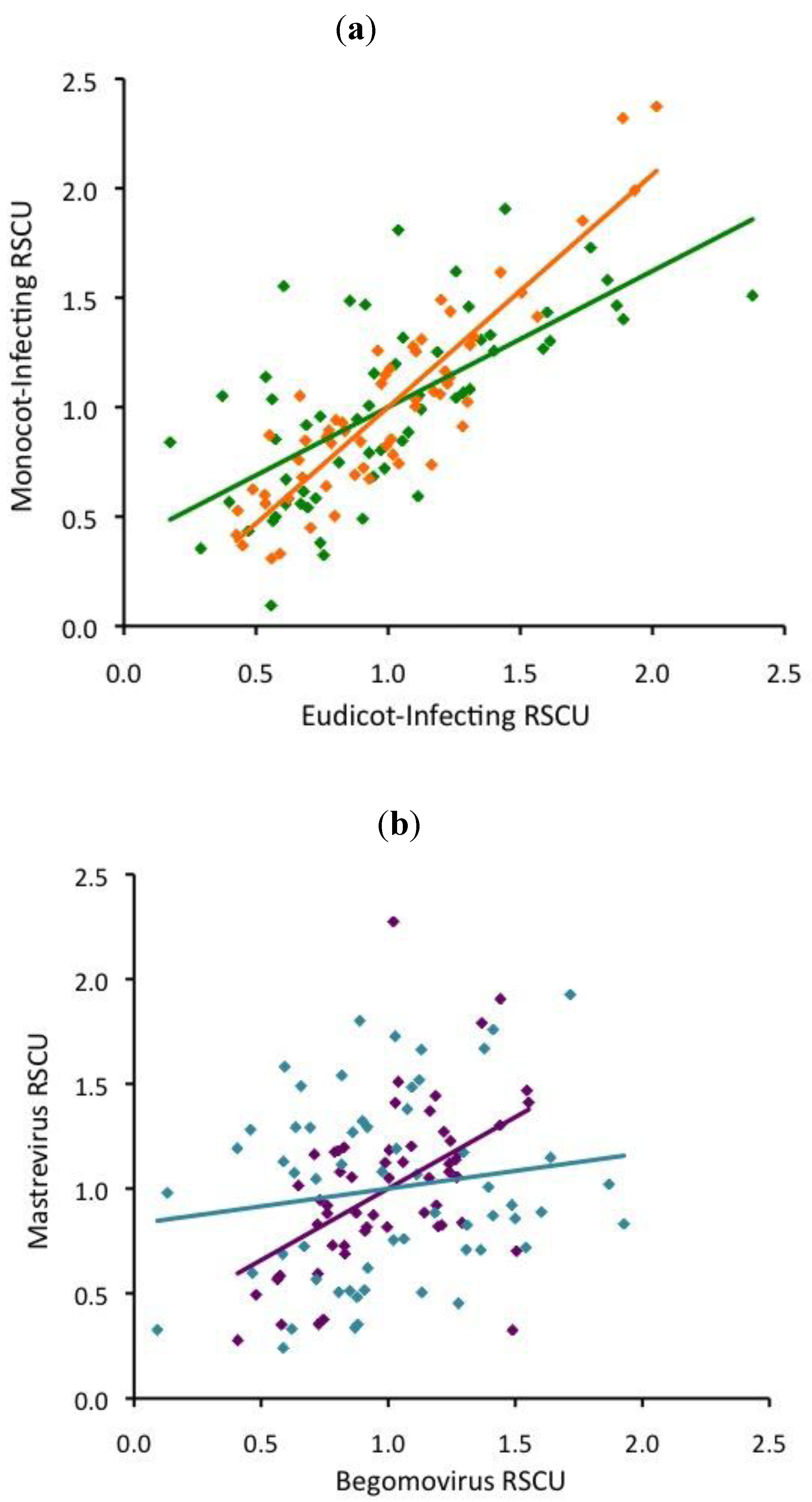
Viruses | Free Full-Text | Base Composition and Translational Selection are Insufficient to Explain Codon Usage Bias in Plant Viruses

Codon usage similarity between viral and some host genes suggests a codon-specific translational regulation - ScienceDirect

Analysis of chromosomes and nucleotides in rice to predict gene expression through codon usage pattern - ScienceDirect
![Natural selection plays a significant role in governing the codon usage bias in the novel SARS-CoV-2 variants of concern (VOC) [PeerJ] Natural selection plays a significant role in governing the codon usage bias in the novel SARS-CoV-2 variants of concern (VOC) [PeerJ]](https://dfzljdn9uc3pi.cloudfront.net/2022/13562/1/fig-3-full.png)
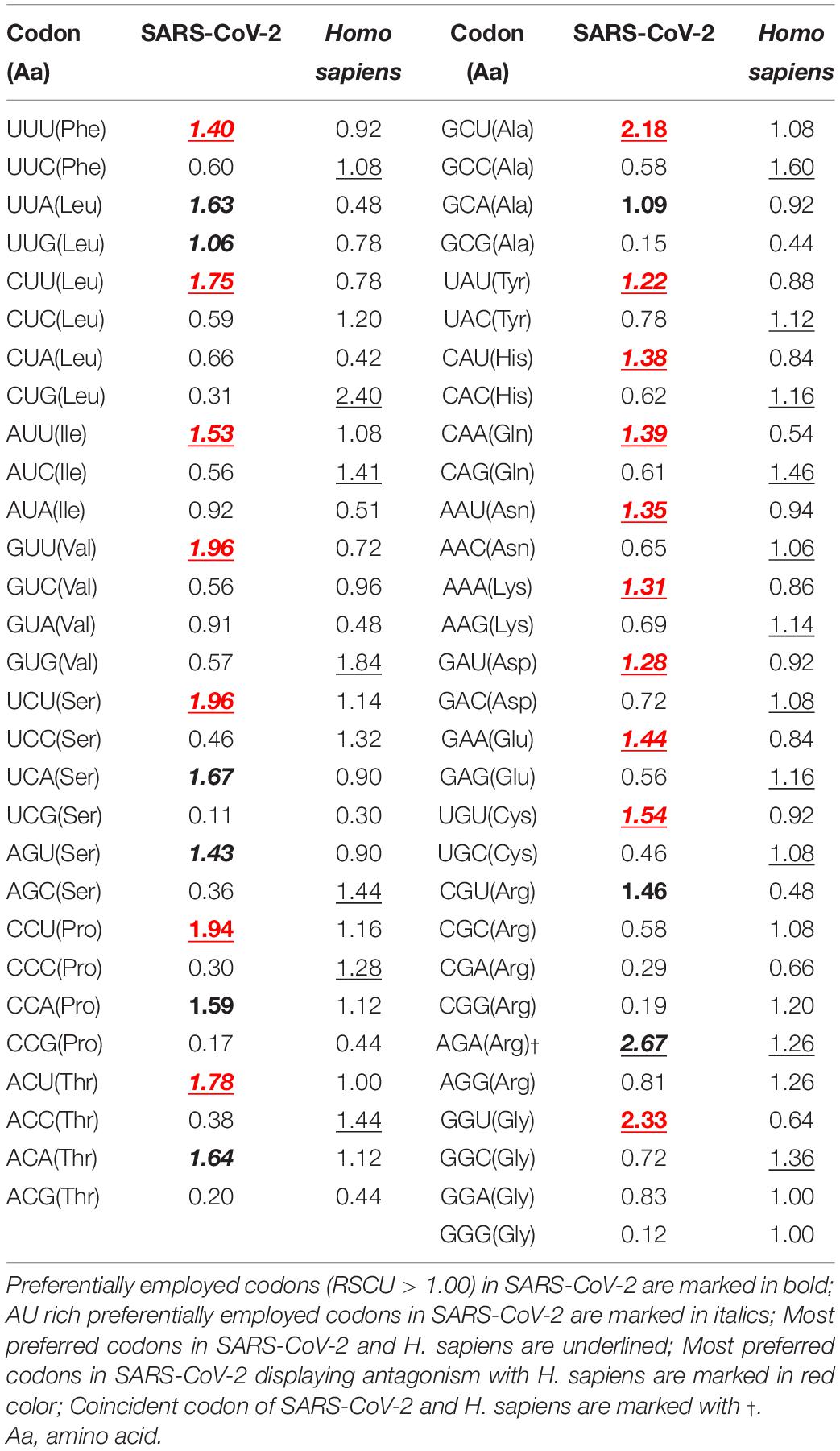
Natural selection plays a significant role in governing the codon usage bias in the novel SARS-CoV-2 variants of concern (VOC) [PeerJ]

Viruses | Free Full-Text | Base Composition and Translational Selection are Insufficient to Explain Codon Usage Bias in Plant Viruses
Correlation between gene expression levels under drought stress and synonymous codon usage in rice plant by in-silico study | PLOS ONE
Codon usage similarity between viral and some host genes suggests a codon-specific translational regulation